PLOS ONE: Comparative Analysis of Membrane Vesicles from Three Piscirickettsia salmonis Isolates Reveals Differences in Vesicle Characteristics

Shift from Gel Based to Gel Free Proteomics to Unlock Unknown Regulatory Network in Plants: A Comprehensive Review
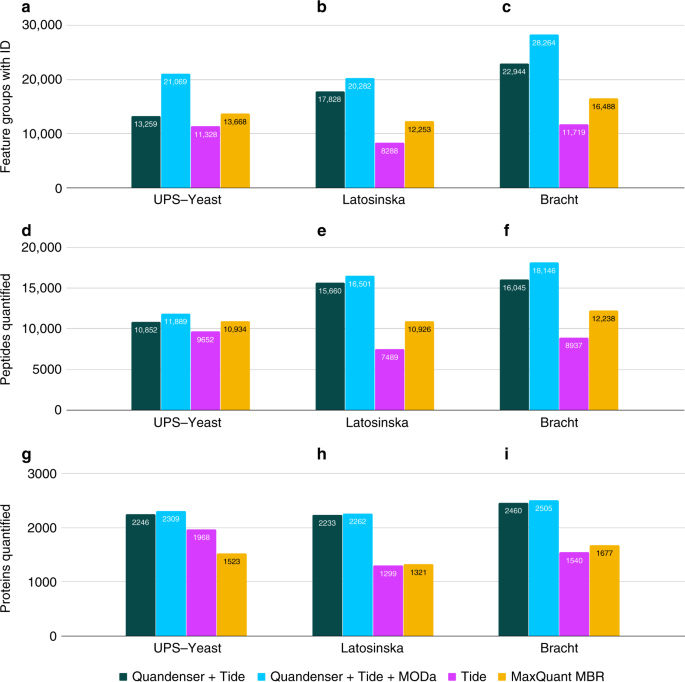
Focus on the spectra that matter by clustering of quantification data in shotgun proteomics | Nature Communications
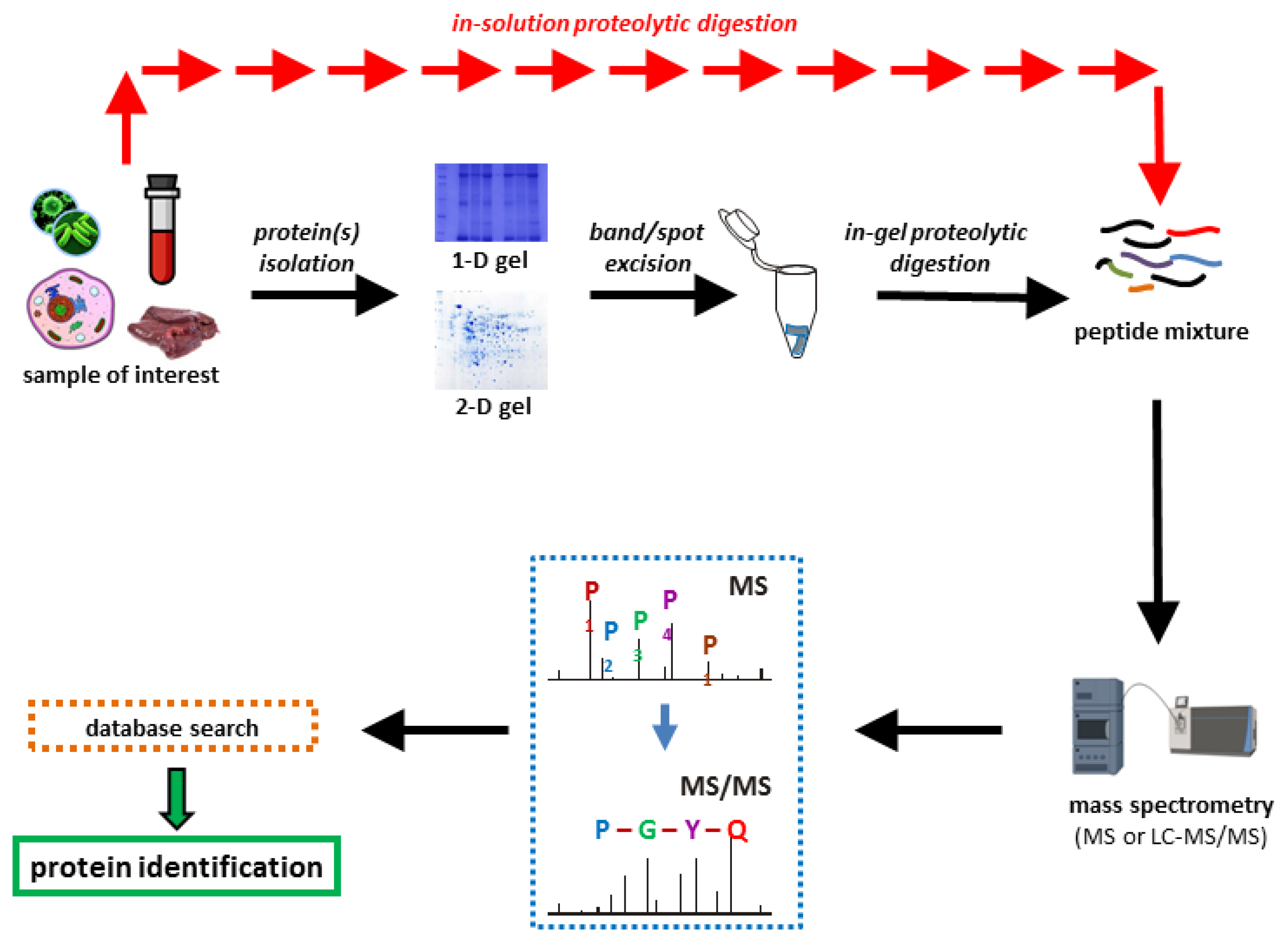
Proteomes | Free Full-Text | A Critical Review of Bottom-Up Proteomics: The Good, the Bad, and the Future of This Field | HTML
Label-free shotgun proteomics and metabolite analysis reveal a significant metabolic shift during citrus fruit development - UNT Digital Library

The suggested three-step workflow of shotgun proteomics analysis that includes contamination/infection detection.

Label‐free deep shotgun proteomics reveals protein dynamics during tomato fruit tissues development - Szymanski - 2017 - The Plant Journal - Wiley Online Library

Significance Analysis of Spectral Count Data in Label-free Shotgun Proteomics - Molecular & Cellular Proteomics

Label free shotgun proteomics for the identification of protein biomarkers for beef tenderness in muscle and plasma of heifers - ScienceDirect

ProtyQuant: Comparing Label-Free Shotgun Proteomics Datasets Using Accumulated Peptide Probabilities | Analytical Chemistry | ChemRxiv | Cambridge Open Engage

Labeling and Label-Free Shotgun Proteomics Quantification in the Research of Cardiovascular Diseases | SpringerLink

ProtyQuant: Comparing label-free shotgun proteomics datasets using accumulated peptide probabilities - ScienceDirect

Targeted Proteomics Guided by Label-free Quantitative Proteome Analysis in Saliva Reveal Transition Signatures from Health to Periodontal Disease* - Molecular & Cellular Proteomics

Quantitative Mass Spectrometry-Based Approaches in Cardiovascular Research | Circulation: Cardiovascular Genetics

Label free shotgun proteomics for the identification of protein biomarkers for beef tenderness in muscle and plasma of heifers - ScienceDirect

Robust Summarization and Inference in Proteome-wide Label-free Quantification - Molecular & Cellular Proteomics









